Cheminformatics on the Web (2021)
Here, I have compiled a list of some libraries and possible ideas. I, personally, like static websites which don't require a server side application and can be hosted on platforms like GitHub Pages. Or, just by opening the HTML file and running it in your browser. WebAssembly (Wasm) has made running code written for other platforms on the web relatively easier. Combine Wasm with some pure JavaScript libraries, and you get a platform to quickly amp up your speed in some common tasks.
RDKit
RDKit bundles a minimal JavaScript Wrapper in their core RDKit suite. This is perfect for generating 2D Figures (HTML5 Canva/SVGs), Canonical SMILES, Descriptors e.t.c
Substructure Matching
This can be used to flag undesirable functional groups in a given compound. Create a simple key:value pairs of name:SMARTS and use it to highlight substructure matches. Thus, something like PostEra's Medicinal Chemistry Alert can be done with RDKit-JS alone.
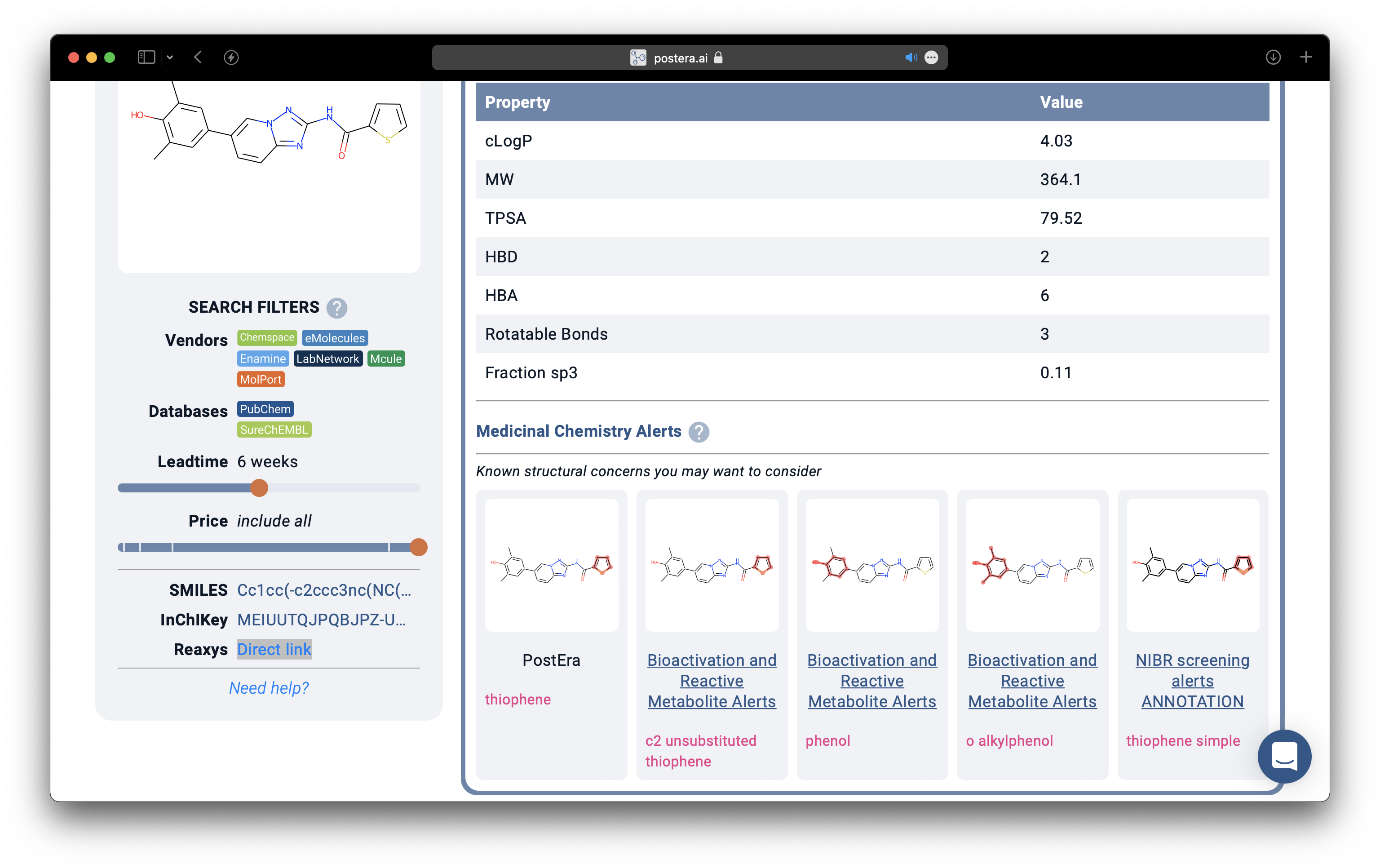
Computing Properties
This is useful to calculate basic properties of a given compound.
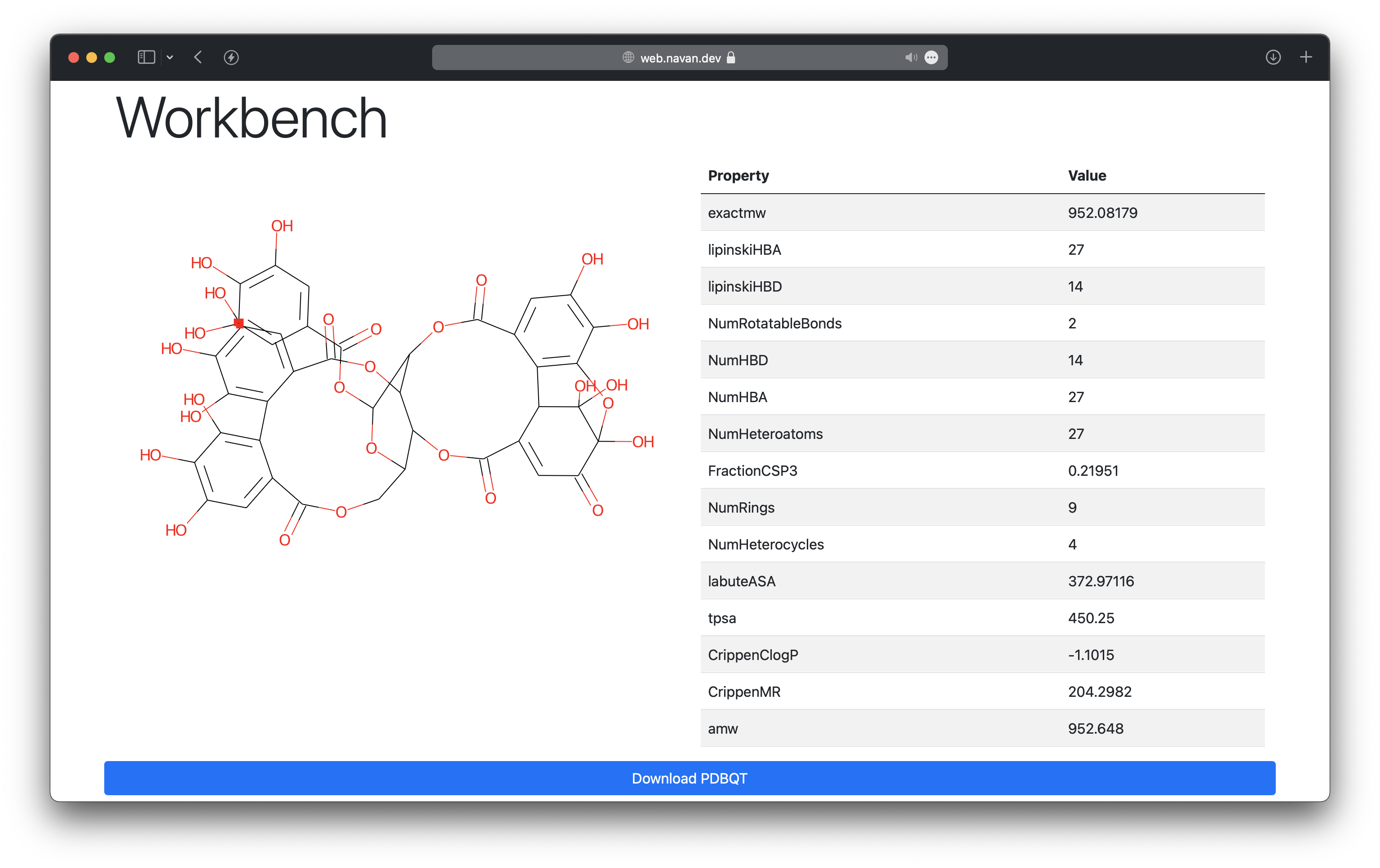
Webina - Molecular Docking
Webina is a JavaScript/Wasm library that runs AutoDock Vina, which can enable you to run Molecular Docking straight in the browser itself.
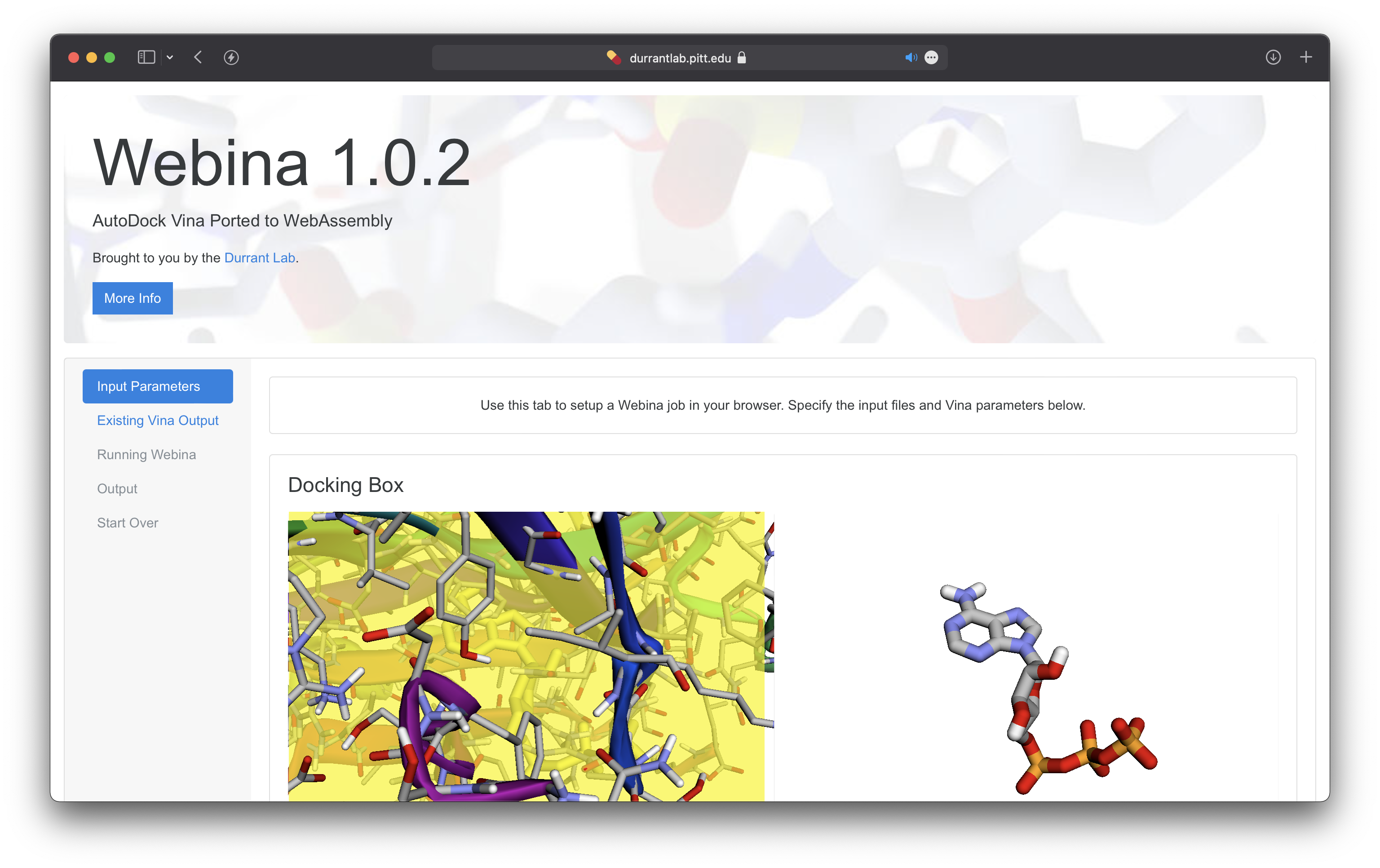
Obviously, it takes a few hits in the time to complete the docking because the code is transpiled from C++ to Wasm. But, the only major drawback (for now) is that it uses SharedArrayBuffer. Due to Spectre, this feature was disabled on all browsers. Currently, only Chromium-based and Firefox browsers have reimplemented and enabled it. Hopefully, soon, this will be again supported by all major browsers.
Machine Learning
Frameworks have now evolved enough to allow exporting models to be able to run them through JavaScript/Wasm backend. An example task can be NER or Named-entity Recognition. It can be used to extract compounds or diseases from a large blob of text and then matched with external references. Another example is target-prediction right in the browser: CHEMBL - Target Prediction in Browser
CHEMBL Group is first training the model using PyTorch (A Python ML Library), then converting it to the ONNX runtime. A model like this can be directly implemented in TensorFlow, and then exported to be able to run with TensorFlow.js
Cheminfo-to-web
The project aims to port cheminformatics libraries into JavaScript via Emscripten. They have ported InChI, Indigo, OpenBabel, and OpenMD
Kekule.js
It is written by @partridgejiang, who is behind the Cheminfo-to-web project
It is molecule-centric, focusing on providing the ability to represent, draw, edit, compare and search molecule structures on web browsers.
Browser Extensions
The previous machine learning examples can be packaged as browser-extensions to perform tasks on the article you are reading. With iOS 15 bringing WebExtensions to iOS/iPadOS, the same browser extension source code can be now used on Desktop and Mobile Phones. You can quickly create an extension to convert PDB codes into links to RCSB, highlight SMILES, highlight output of NER models, e.t.c
Conclusion
I have not even touched all the bases of cheminformatics for the web here. There is still a lot more to unpack. Hopefully, this encourages you to explore the world of cheminformatics on the web.

